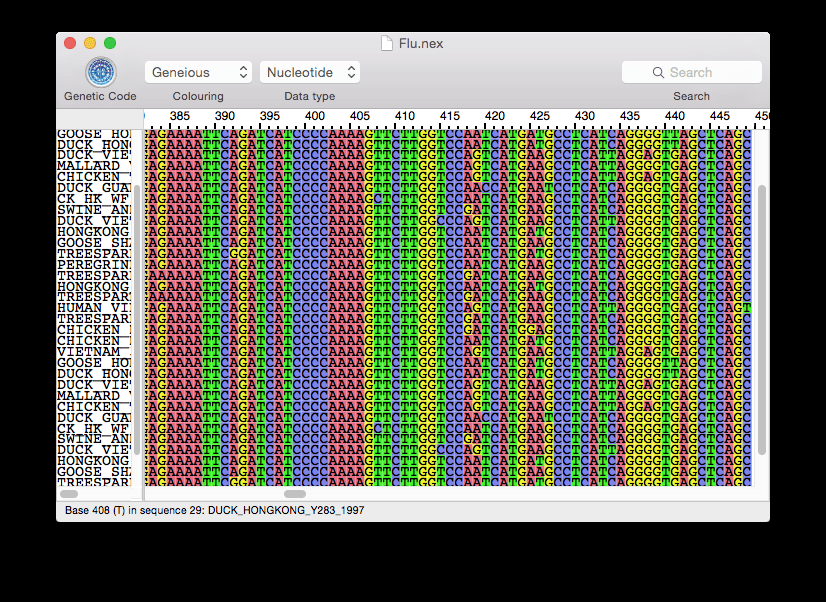
Fixed: Calculation of consensus sequence in read mappings: Sometimes a majority of gaps would be ignored and a base erroneously introduced in the consensus sequence.The annotation was only displayed on the first line. Fixed: Annotations spanning the sequence from start to end did not display right when the sequence was wrapped.Exporting fastq format no longer includes redundant name of the read in the quality score line.They always applied to the first sequence although the fragment was located on another sequence (as indicated in the table). Fixed: When using multiple template sequences, the choices to open or annotate a fragment from the fragment table did not work properly.a T would still be a mismatch if the template sequence has an R, which means either A or G). Note that the primer base of course need to be covered by the ambiguity symbol (e.g. If your template sequence contains ambiguity nucleotides (like N, Y etc), these will no longer count as mismatches when checking your primers.Find Binding Sites and Create Fragments improved:.You can search for barcodes (MIDs) on both strands, supporting new 454 protocol.A summary report is now available with an overview of the number of reads per bar code.The existing Gateway Cloning tool has been expanded so that you can easily recombine several fragments as well as continue using it for the standard Gateway Cloning. You can perform multi-site gateway cloning and in a few clicks create your expression clones with multiple fragments. It also includes a migration to version 68 of Ensembl a new variation dataset for Puccinia graminis a structural variation dataset for Sorghum bicolor imported from the database of genomic variants archive and improvements to assembly, annotation and variation datasets among other updates. This release adds two new genomes to Ensembl Fungi - Leptosphaeria maculans and Trichoderma virens - bringing the total genomes supported to 354. Also in this version, the water orientation VIDA extension has been completely rewritten to make it easier to use and to find key waters and understand their interactions and several bugs have been fixed. It also produces hypotheses of ligand modifications to improve affinity, based on the energetics of the water environment directly adjacent to the ligand. The release includes a new command-line program called GamePlan that runs several SZMAP calculations and analyzes the results to examine how the existing ligand chemistry aligns with the pocket environment. OpenEye Scientific Software has released SZMAP v1.1.0, the latest version of the company’s software for studying water’s role in molecular interactions. It also improves the assembly of data from Ion Torrent’s PGM and adds a new coverage normalization capability to NextGene’s Floton assembler generates reports automatically and imports and filters data from the Catalogue of Somatic Mutations in Cancer among other updates. The release includes new tools for HLA analysis and accepts the SOLiD XSQ data format. 
SoftGenetics has also released a new version of its NextGene software, which is used for analyzing next-generation sequence data. Users also have access to applications in the GeneMarker software suite, such as multiplex ligation-dependent probe amplification, trisomy analysis, and more. SoftGenetics has released GeneMarkerMTP, a multi-template processor for use in genotyping applications.Īccording to its developers, GeneMarkerMTP can process varying chemistries or size standards in a single analysis run. Additionally, users can create workflows that combine various tools from the toolbox into a single analysis step have access to tools for advanced variant detection, which are well suited for genomes of higher ploidy and can easily download genomic sequences and annotation data from public databases among other capabilities. This week, CLC Bio released CLC Genomics Workbench 5.5, the latest version of its desktop software and CLC Genomics Server 4.5, the newest release of its enterprise software.īoth software packages now include tools to build complete resequencing workflows including read mapping, variant detection, and downstream analysis. The latest version of the tool can accept and publish very large images, made up of many individual tiles, and make them viewable online. 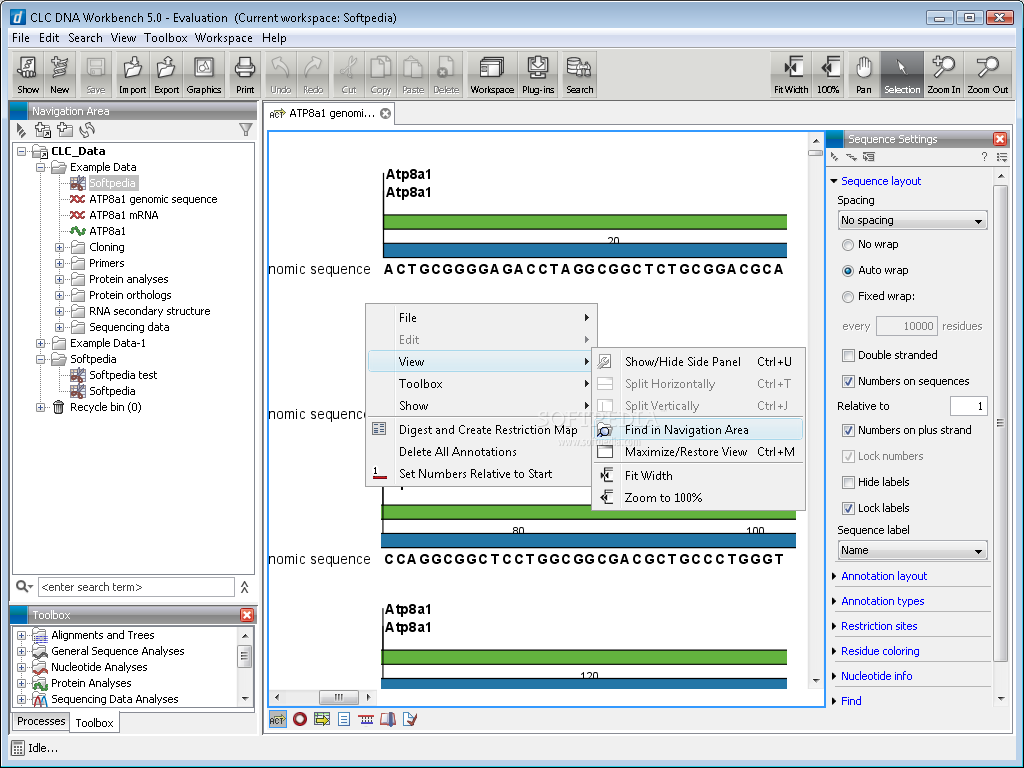
Glencoe Software and the Journal of Cell Biology have updated the JCB DataViewer, a platform for presenting, sharing, and archiving published scientific image data that is based on technology developed by the Open Microscopy Environment. Advances in Clinical Genomics Profiling.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
